-Search query
-Search result
Showing 1 - 50 of 336 items for (author: simon & ae)

EMDB-19406: 
Structure of the human DDB1-DDA1-DCAF15 E3 ubiquitin ligase bound to compound furan 12
Method: single particle / : Shilliday F, Lucas SCC, Richter M, Michaelides IN, Fusani L

EMDB-19407: 
Structure of the human DDB1-DDA1-DCAF15 E3 ubiquitin ligase bound to compound furan 24
Method: single particle / : Shilliday F, Lucas SCC, Richter M, Michaelides IN, Fusani L

PDB-8rox: 
Structure of the human DDB1-DDA1-DCAF15 E3 ubiquitin ligase bound to compound furan 12
Method: single particle / : Shilliday F, Lucas SCC, Richter M, Michaelides IN, Fusani L

PDB-8roy: 
Structure of the human DDB1-DDA1-DCAF15 E3 ubiquitin ligase bound to compound furan 24
Method: single particle / : Shilliday F, Lucas SCC, Richter M, Michaelides IN, Fusani L

EMDB-40603: 
GPR161 Gs heterotrimer
Method: single particle / : Hoppe N, Manglik A, Harrison S

PDB-8smv: 
GPR161 Gs heterotrimer
Method: single particle / : Hoppe N, Manglik A, Harrison S
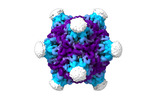
EMDB-19024: 
Structure of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG
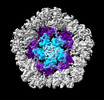
EMDB-19025: 
Structure of the five-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG
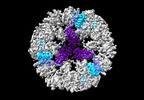
EMDB-19026: 
Structure of the three-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG
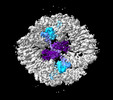
EMDB-19027: 
Structure of the two-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb3: 
Structure of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb4: 
Structure of the five-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb5: 
Structure of the three-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb7: 
Structure of the two-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

EMDB-16976: 
Human tRNA guanine transglycosylase (TGT) bound to tRNAAsp
Method: single particle / : Sievers K, Neumann P, Susac L, Trowitzsch S, Tampe R, Ficner R

PDB-8omr: 
Human tRNA guanine transglycosylase (TGT) bound to tRNAAsp
Method: single particle / : Sievers K, Neumann P, Susac L, Trowitzsch S, Tampe R, Ficner R

EMDB-16805: 
Cryo-EM structure of PcrV/Fab(30-B8)
Method: single particle / : Yuan B, Simonis A, Marlovits TC

EMDB-16807: 
Cryo-EM structure of PcrV/Fab(11-E5)
Method: single particle / : Yuan B, Simonis A, Marlovits TC

PDB-8cr9: 
Cryo-EM structure of PcrV/Fab(30-B8)
Method: single particle / : Yuan B, Simonis A, Marlovits TC

PDB-8crb: 
Cryo-EM structure of PcrV/Fab(11-E5)
Method: single particle / : Yuan B, Simonis A, Marlovits TC

EMDB-17787: 
4.0 angstrom map of outward-facing MFS transporter MHAS2168, a homologue of M. tuberculosis Rv1410, in complex with a megabody
Method: single particle / : Remm S, Gonda I, Seeger MA

EMDB-40334: 
Human OCT1 (Apo) in inward-open conformation
Method: single particle / : Zeng YC, Sobti M, Stewart AG

EMDB-40335: 
Human OCT1 bound to diltiazem in inward-open conformation
Method: single particle / : Zeng YC, Sobti M, Stewart AG

EMDB-40336: 
Human OCT1 bound to fenoterol in inward-open conformation
Method: single particle / : Zeng YC, Sobti M, Stewart AG

EMDB-40337: 
Human OCT1 bound to metformin in inward-open conformation
Method: single particle / : Zeng YC, Sobti M, Stewart AG

EMDB-40339: 
Human OCT1 bound to thiamine in inward-open conformation
Method: single particle / : Zeng YC, Sobti M, Stewart AG

PDB-8sc1: 
Human OCT1 (Apo) in inward-open conformation
Method: single particle / : Zeng YC, Sobti M, Stewart AG

PDB-8sc2: 
Human OCT1 bound to diltiazem in inward-open conformation
Method: single particle / : Zeng YC, Sobti M, Stewart AG

PDB-8sc3: 
Human OCT1 bound to fenoterol in inward-open conformation
Method: single particle / : Zeng YC, Sobti M, Stewart AG

PDB-8sc4: 
Human OCT1 bound to metformin in inward-open conformation
Method: single particle / : Zeng YC, Sobti M, Stewart AG

PDB-8sc6: 
Human OCT1 bound to thiamine in inward-open conformation
Method: single particle / : Zeng YC, Sobti M, Stewart AG

EMDB-18383: 
Cryo-EM structure of SidH from Legionella pneumophila
Method: single particle / : Sharma R, Weis F, Bhogaraju S

EMDB-18407: 
Cryo-EM structure of SidH from Legionella pneumophila in complex with LubX
Method: single particle / : Sharma R, Adams M, Bhogaraju S

PDB-8qfs: 
Cryo-EM structure of SidH from Legionella pneumophila
Method: single particle / : Sharma R, Weis F, Bhogaraju S

PDB-8qhc: 
Cryo-EM structure of SidH from Legionella pneumophila in complex with LubX
Method: single particle / : Sharma R, Adams M, Bhogaraju S
EMDB-28013: 
Cryo-EM structure of the Glutaminase C core filament (fGAC)
Method: single particle / : Ambrosio AL, Dias SM, Quesnay JE, Portugal RV, Cassago A, van Heel MG, Islam Z, Rodrigues CT

PDB-8ec6: 
Cryo-EM structure of the Glutaminase C core filament (fGAC)
Method: single particle / : Ambrosio AL, Dias SM, Quesnay JE, Portugal RV, Cassago A, van Heel MG, Islam Z, Rodrigues CT

EMDB-40181: 
Human TRPV3 tetramer structure, closed conformation
Method: single particle / : Lansky S, Betancourt JM, Scheuring S

EMDB-40183: 
Human TRPV3 pentamer structure
Method: single particle / : Lansky S, Betancourt JM, Scheuring S

PDB-8gka: 
Human TRPV3 tetramer structure, closed conformation
Method: single particle / : Lansky S, Betancourt JM, Scheuring S

PDB-8gkg: 
Human TRPV3 pentamer structure
Method: single particle / : Lansky S, Betancourt JM, Scheuring S

EMDB-40789: 
BAP1/ASXL1 bound to the H2AK119Ub Nucleosome
Method: single particle / : Thomas JF, Valencia-Sanchez MI, Armache KJ

EMDB-40790: 
Map focused on acidic patch BAP1/ASXL1 bound to the H2AK119Ub Nucleosome
Method: single particle / : Thomas JF, Valencia-Sanchez MI

EMDB-40791: 
Overall map of BAP1/ASXL1 bound to the H2AK119Ub Nucleosome
Method: single particle / : Thomas JF, Valencia-Sanchez MI

PDB-8svf: 
BAP1/ASXL1 bound to the H2AK119Ub Nucleosome
Method: single particle / : Thomas JF, Valencia-Sanchez MI, Armache KJ
EMDB-17944: 
Cryo-EM structure of an asymmetric nucleosome (10-N-0) from a Glacios/Falcon3EC
Method: single particle / : Gaullier G

EMDB-40184: 
Structure of the Spizellomyces punctatus Fanzor (SpuFz) in complex with omega RNA and target DNA
Method: single particle / : Xu P, Saito M, Zhang F

PDB-8gkh: 
Structure of the Spizellomyces punctatus Fanzor (SpuFz) in complex with omega RNA and target DNA
Method: single particle / : Xu P, Saito M, Zhang F

EMDB-17114: 
CAND1-CUL1-RBX1-SKP1-SKP2-CKS1-CDK2
Method: single particle / : Shaaban M, Clapperton JA, Ding S, Maeots ME, Enchev RI, Maslen S, Skehel JM

EMDB-17115: 
CAND1-CUL1-RBX1-DCNL1
Method: single particle / : Shaaban M, Clapperton JA, Ding S, Maeots ME, Enchev RI, Maslen SL, Skehel JM
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
